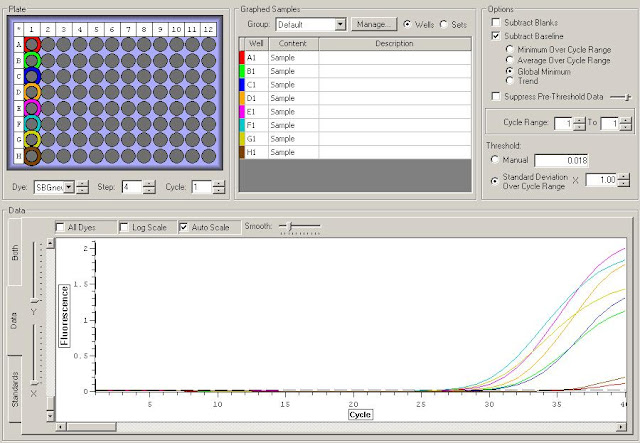
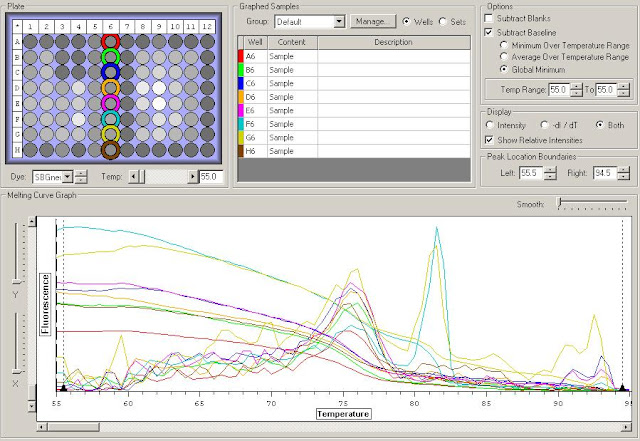
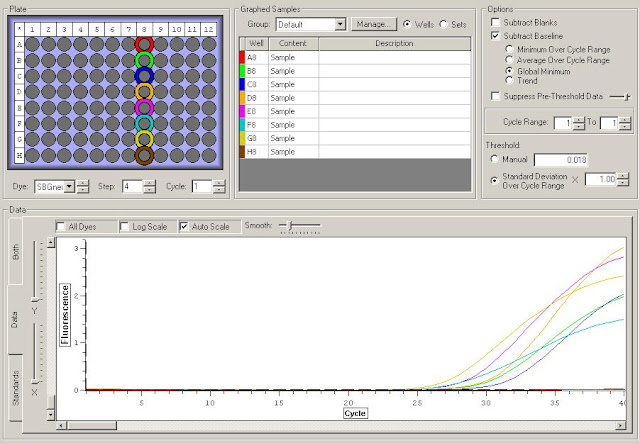
First I need to figure out where in the sequence does the original primer amplify. To do this I have to upload the original sequence and the primer sequences into NCBI. For this I'm looking at the TLR2.1 primer since it has the most interesting results from the primer check.
The original primer sequences for TLR2.1:
FWD ACAAAGATTCCACCCGGCAA
REV ACACCAACGACAGGAAGTGG
Product Size 109 bp
The original sequence for target gene:
tcaaaacagattttggagtgtaacgtctttaaaaattactaatacagcaacaatgaaatcattttgccatgaattgcttcacgatttgccagaacagcttctggccatttctattcattgtccactctacatgagatgctgacttgagaagcaccttcagagattgattaatatgttcagaacgaatatcttgtaatatgatcagcaatagcttgtcaccgcctccgtcagccaaagtgtggttggcaatggcggcttcgtacttacaccattgatcgtccaagaaattattggacagcaccaaaatgaattttctgctgacttcaatgctttccagaaagtcatccacaaagattccacccggcaagatatctctctcgtgcaaacaaagcttgtacttgtttctctctaccatctgaacaagttcagacattatccacttcctgtcgttggtgttgtaagcaacaaaaccatcatagaggaattcatcatctgtaaatcttttataccccatgggttttctgtttttgcatgtgtatatataatacttaatgtcccaacgaaactttctcaacagaacaatggtcaaaatcaccaaggacactaatgcaaacgaagcgcatactagaatgacataggggcttattttgtgacattcggacgcaggatcaaagtgtgttatactttttcctttcaagtctgacggggaagcacagatgtattgatgaggatatccaacagtcctattttggttgttttcaacccagagtcggaaccattccagttggcatccacagtccaatggattcaaggacacatcaattcgaaaatcttttttcttccaaaggaaagcaggcagagaagattcattgatactgctcaaacgattgccacgaaaaatcaattcattcaatctagtcaaattttgaacaccgctctttcgaatttctgttattcggttaacactgtagtcaacggacgttagctgtgaaaaatcagtcagtgtagtcatttgatcgatttcgccattaacaatagccaaaactgacagatcctttagaggtgatagcatctctgcaacatcagtgttggatagatcggtgttcattattttcaatgcttttatatttctacattcagaaaagatgtatttgaagcctcctacgcttttttccaccatacgtccgatagagagtgttcggagggaattggatgataatgacctaccatctaaagcgactccatgcaagttatcaatgatcaaagtttgcaaacgattcatattcaaaaaggaatacaaggaaatgtcatgaattgatgtgtgtgaaacattaaaaaattcaagggatttcaggcagtagccctcttgtatgaaatttgttgtttcccgaataccattcactgccaggtgcaattgttgcaagtttgggaaggcggctatacgtttctcattggataaacaaaactcgggaatcttctcaaaagaattgttagtaaaatacagttcttttaaagaaggaactgagctatttggcaatttataccttgagattaggttgcttcctaaatcgattttgtggatattgattaatttagcgaaggatttaaaag
First I run the sequence and the primers through NCBI Primer Blast:
Once the Primer Blast returns the primer target. I can then find where the primer begins and ends in the original sequence.
The forward primer begins at roughly 340 bp and the reverse primer begins at roughly 460 bp. Using these numbers I subtract 100 bp from the start of the forward primer and add 100 bp to the beginning of the reverse primer. This extends the range to 240 bp - 560 bp which encompasses the original target and primer area as well as 100 bp on either side. To give the Primer Blast some leeway to produce the new primers, I give a 100 bp range for the forward and reverse primers. The range for the forward primer becomes 140-240 bp. The range for the reverse primer becomes 560-660 bp. I also required the product size to be between 325 bp and 1000 bp long. This is so that the 100 bp on either side plus the 109 bp original target are amplified.
The primers that NCBI now covers the entire area that needs to be sequenced. All I have to do is choose primers that have low self complimentarity.
After looking at the offered primers I chose one that should give a 425 bp product with low self complimentarity.
This is the quickest way I've found to take the primers I produced originally and create primers to sequence the target area of interest. If this works I'll do this with all the primers of interest that worked in the previous primer runs.